Cassie S. Mitchell, Ph.D.

Assistant Research Professor
Biomedical Engineering
Georgia Institute of Technology and Emory University School of Medicine
Believe in yourself, take risks, and never give up
On my desk is displayed a sign that says, “Never, never, never give up.” It serves as my motto and personal reminder of the importance of each day’s work towards my ultimate goal of alleviating neurological disease. As a C5-6 wheelchair-bound quadriplegic from the neurological disease Neuromyelitis Optica, I have an intimate understanding of the impact of neuropathologies and a passion to leverage my personal and scientific research experiences to positively impact the lives of patients.
I obtained my Ph.D. in biomedical engineering at Georgia Institute of Technology and Emory University School of Medicine. My background was in computational modeling, so it was important to me that I diversify my skills and try the “wet” side of science. During my pre-thesis rotations, I worked on in vivo, in vitro, and in silico neuroscience. Thankfully, my natural ability leaned in the theoretical direction, because my physical disability made traditional “wet lab” experimental research very difficult. The need to tape instruments in your hand to perform procedures gives new meaning to “standard experimental error!”.
Nonetheless, the experience I obtained in my rotations remains highly valuable, as it enabled me to effectively collaborate with researchers and understand data obtained from those fields. In fact, the utility of my current computational research depends on my understanding of experimental biology. A computer simulation won’t tell you if an experiment is actually feasible in a lab, so I have to stay up-to-date on the latest experimental methodologies, techniques, and equipment, in order to make sure that the predictions I make will be useful to the field.
During my graduate work, my research focused on developing new methods to analyze pathophysiology. I realized during that time that, despite all the emphasis on innovative, trans-disciplinary research, it is actually very hard to bridge the gap between the theoretical, experimental, and clinical domains. From my perspective, communication between these three fields was broken and I wanted my future research to bridge that gap.
During my postdoc, I began to search for funding opportunities that fit my career goals. The Faculty K01 Career Award appealed to me because you could craft a plan customized to the skills you had and the ones you wanted to gain. I developed a proposal to integrate experimental and clinical data to produce clinically meaningful, predictive outcomes in amyotrophic lateral sclerosis (ALS). In crafting my application, I focused on what I would need to successfully integrate across disparate disciplines. I identified as mentors an experimental neurophysiologist, a clinical neurologist, and a statistician, who I felt would give me the diversity of perspective and thought that I would need to succeed.
The Faculty K01 Award has dramatically and positively shaped my career and has directly enabled my success to this day. I spent the first two years doing intense clinical training (learning patient interactions, survey and medical records structure, and IRB protocol development) while laying the research foundation for my lab, which included developing data paths for three different informatics projects.
Early on, I realized that in order to do my research at the scale I wanted, I needed more people in the lab. Therefore, during years two and three of the K01, I methodically expanded my lab from three people to over fifty, developing the “undergraduate research corporation” model, which allows the staff to pursue informatics projects of unprecedented size while providing undergraduates with technical and management opportunities customized for their interests and skills.
Many of my colleagues initially doubted the productivity of having such a large team of undergraduates to manage. They encouraged me to scale down my research and wait for the data collection and processing I needed to become automated. I took their concerns to heart, but ultimately felt that the benefits outweighed the risks. Fortunately, with a lot of patience and hard work, the risk paid off. In the last four years, my lab has collected more than 1,000,000 data points to analyze—work that would have swamped the bandwidth of most labs. These data allowed me to apply for and receive an R21 and an R03 grant during my K01.
When not fighting neurological disease via research or mentoring, I am racing it as an elite athlete on the USA Paralympic Track & Field team. I competed in the London 2012 Games, and I am currently training for the 2016 Rio de Janeiro Olympics & Paralympics. I’ve found that athletics facilitates my creativity; my best ideas come when working on a training session. It allows me space to decompress, and my research has benefited because it keeps me fresh and rejuvenates me. I’ve been asked how I find the time to compete at this level, and my best advice is to prioritize—in all aspects of your life, research, hobbies, family, etc.—the things that you are really passionate about. When it comes to research, do not chase fads, change yourself, or walk away from your research passion simply to perceptually increase odds of funding—and live by the motto: Never, never, never give up!
Current Research
My research focuses on the development of methodologies to better assess and treat devastating neuropathologies. I describe myself as a "pathology forecaster" who uses published data to quantitatively predict disease mechanisms and treatment outcomes. Much like the weather forecasters who utilize computer models that track severe weather to warn communities, my lab’s novel predictive disease analyses expedite experimental research and clinical trials by identifying the most likely disease causes and the most promising treatments, so we can get to the right solutions faster.
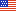
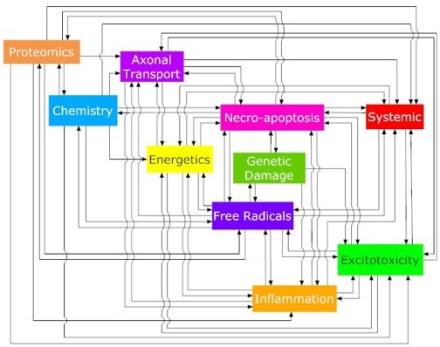
One such project involves “dynamic meta-analysis” (DMA), a technique developed by my lab that incorporates time and interactions between factors. We applied DMA to a database of every quantifiable data point (300,000+) in the literature on a mouse model of amyotrophic lateral sclerosis. This produced a model system that is trying to re-stabilize but overcompensates, oscillating above and below homeostasis. Using pathology dynamics, we were able to partially re-stabilize the system by applying combination treatments targeting three or more processes known to be involved or affected by the disease, thus providing predictions for novel treatments.